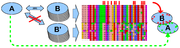
Many protein families involved in protein-protein interaction (PPI) contain sub-families that interact with different protein binding partners. We have applied the Sequence Harmony method (SH), which is able to detect specificity sites from a multiple sequence alignment (MSA) containing sub-families. The input was a dataset of MSAs of interacting protein families, each time containing a set of non-interacting paralogous sequences. Exploiting the differences in sequence conservation between the binding and non-binding groups by \SH, we demonstrate that predicted specificity residues turn out to reside on the protein surface. We also show that we can select interface residues with approximately 14% coverage (true-positive rate) at 27% error (false-positive rate).
The full article is available behind this button:
![]()